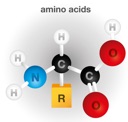
 Understanding the cellular roles of DNA and protein is challenging because they are, in important and meaningful ways, not analogous to anything in the macroscopic world. Proteins are a fascinating study: while cars, watches and houses are built by specialized builders, a single master factory, the ribosome, strings together all cellular proteins. And in a richly meaningful sense, that string then assembles itself (folds) into its final form. And the machines of the cell are almost unimaginably diverse in their forms and functions, the latter of which derive once again from within: the amino acid building blocks and precisely where they come to rest in the final three dimensional structure.
Understanding the cellular roles of DNA and protein is challenging because they are, in important and meaningful ways, not analogous to anything in the macroscopic world. Proteins are a fascinating study: while cars, watches and houses are built by specialized builders, a single master factory, the ribosome, strings together all cellular proteins. And in a richly meaningful sense, that string then assembles itself (folds) into its final form. And the machines of the cell are almost unimaginably diverse in their forms and functions, the latter of which derive once again from within: the amino acid building blocks and precisely where they come to rest in the final three dimensional structure.
All posts by Bruce - 3. page
I've taught introductory biology for... a long time now--at U. of Arizona, Princeton, Stockton (NJ) and Sarah Lawrence College, as well as online (Wyzant). Currently I'm retired; all the teaching I do is for the love of it.
Plea for pedigrees: pedigree deduction in Introductory Biology
 Pedigree-solving can be perceived as passé in an era in which every single nucleotide of an individual’s genome can be learned (relatively) affordably, parents be damned. But I don’t think that’s the value in teaching pedigree deduction; indeed, I question whether it ever should have been. As with many exercises in Introductory Biology, I think we need to make careful distinctions between means and ends. Pedigree deduction can be employed as one of a class of near-perfect problem-solving opportunities, employing limited, easy-to-grasp tools, reflecting core biology (meiosis, randomness), and representing an unforgiving series of clear deductions. To my mind, the question should be how do we make sure these aspects are represented in tasks and assessments involving pedigrees.
Pedigree-solving can be perceived as passé in an era in which every single nucleotide of an individual’s genome can be learned (relatively) affordably, parents be damned. But I don’t think that’s the value in teaching pedigree deduction; indeed, I question whether it ever should have been. As with many exercises in Introductory Biology, I think we need to make careful distinctions between means and ends. Pedigree deduction can be employed as one of a class of near-perfect problem-solving opportunities, employing limited, easy-to-grasp tools, reflecting core biology (meiosis, randomness), and representing an unforgiving series of clear deductions. To my mind, the question should be how do we make sure these aspects are represented in tasks and assessments involving pedigrees.
Dependencies and foundations: introductory biology design
 We’ve long since passed the days when a strategy of “Just teach the students everything in the field” even begins to define a good introductory biology (or even specialized biology) course. Alas, this rut has ensnared textbooks; even those that start out nobly slender quickly bulk up to monstrous size. A related problem is the question of topic order–I think the historical approach (generally all the chemistry, a difficult to comprehend overview, then a firehose of detail) is out of touch with modern thinking about learning. As a way out of the maze, I suggest identifying the Big Ideas, selecting powerful, highly interlinked exemplars (hemoglobin profiled in this role here) and keeping a careful eye on what must come before what. In this post, I’d like to present the ideas of dependency (what must be understood prior to introducing a topic) and foundation (what understandings are enabled once the topic is mastered).
We’ve long since passed the days when a strategy of “Just teach the students everything in the field” even begins to define a good introductory biology (or even specialized biology) course. Alas, this rut has ensnared textbooks; even those that start out nobly slender quickly bulk up to monstrous size. A related problem is the question of topic order–I think the historical approach (generally all the chemistry, a difficult to comprehend overview, then a firehose of detail) is out of touch with modern thinking about learning. As a way out of the maze, I suggest identifying the Big Ideas, selecting powerful, highly interlinked exemplars (hemoglobin profiled in this role here) and keeping a careful eye on what must come before what. In this post, I’d like to present the ideas of dependency (what must be understood prior to introducing a topic) and foundation (what understandings are enabled once the topic is mastered).
Great experiments: Drosophila development in one fell swoop
 A Great Experiment should have several qualities: it should address a Big Question of its time, it should ‘move the ball’ a good ways and be universally seen to do so, it should entail an approach that is insightful and hopefully clever, and should represent a careful foundation of what system and techniques should be employed–or invented. “Mutations affecting segment number and polarity in Drosophila” by Christiane Nüsslein-Volhard and Eric Wieschaus is just such a paper. FYI, the Lasker Award folks and the Nobel Prize selection committee thought so too. It represents a combination of a huge question (where does multicellular form come from given a single-celled start?), brilliant recognition of ideal experimental system (Drosophila combines ancient genetic tools with a ‘body plan’ that is scribed on the surface of the early embryo), invention (I believe) of Drosophila ‘replica plating’ and resulted in a series of mutants that by their existence and phenotypes left us with a clear idea of how the body plan of Drosophila is laid out through sequential gene-expression driven subdivision into smaller regions. Continue reading
A Great Experiment should have several qualities: it should address a Big Question of its time, it should ‘move the ball’ a good ways and be universally seen to do so, it should entail an approach that is insightful and hopefully clever, and should represent a careful foundation of what system and techniques should be employed–or invented. “Mutations affecting segment number and polarity in Drosophila” by Christiane Nüsslein-Volhard and Eric Wieschaus is just such a paper. FYI, the Lasker Award folks and the Nobel Prize selection committee thought so too. It represents a combination of a huge question (where does multicellular form come from given a single-celled start?), brilliant recognition of ideal experimental system (Drosophila combines ancient genetic tools with a ‘body plan’ that is scribed on the surface of the early embryo), invention (I believe) of Drosophila ‘replica plating’ and resulted in a series of mutants that by their existence and phenotypes left us with a clear idea of how the body plan of Drosophila is laid out through sequential gene-expression driven subdivision into smaller regions. Continue reading
If I were Howard: improving upon the Phage Hunters project
 Undergraduate labs are notoriously dismal, certainly from the student point of view and often from faculty’s as well. One obvious reason is that the design and delivery must to some extent be cookie-cutter because materials and set-up must be planned for what may be dozens of sections, delivered to hundreds of students, and most often constrained to once-a-week, two- or three-hour meetings. The Howard Hughes Medical institute (HHMI) has sponsored the HHMI Phage Hunters (SEA-phage) project (and related UT Austin Freshman Research Initiative) with the goal of addressing several shortcomings of classical Intro Bio labs. I believe that it’s on the verge of being something truly awesome (at least from the point of view of a structure/function discovery-driven instructor) and greatness for ‘my’ target group could be by following the core thinking to different ends.
Undergraduate labs are notoriously dismal, certainly from the student point of view and often from faculty’s as well. One obvious reason is that the design and delivery must to some extent be cookie-cutter because materials and set-up must be planned for what may be dozens of sections, delivered to hundreds of students, and most often constrained to once-a-week, two- or three-hour meetings. The Howard Hughes Medical institute (HHMI) has sponsored the HHMI Phage Hunters (SEA-phage) project (and related UT Austin Freshman Research Initiative) with the goal of addressing several shortcomings of classical Intro Bio labs. I believe that it’s on the verge of being something truly awesome (at least from the point of view of a structure/function discovery-driven instructor) and greatness for ‘my’ target group could be by following the core thinking to different ends.
A mechanistic view of the cell: the glory of because
 Introductory biology can be overwhelming; what we consider to be ‘essential foundation’ could take up an entire undergraduate education. This issue is compounded when we treat the body of knowledge as a collection of names and facts (or when students achieve this view despite our efforts). By viewing the cell as a functioning machine, divided into compartments filled tasks performed by mindless but efficient mechanisms, a course can focus on understanding. A natural framework for student study and learning arises: the mechanics of the cell
Introductory biology can be overwhelming; what we consider to be ‘essential foundation’ could take up an entire undergraduate education. This issue is compounded when we treat the body of knowledge as a collection of names and facts (or when students achieve this view despite our efforts). By viewing the cell as a functioning machine, divided into compartments filled tasks performed by mindless but efficient mechanisms, a course can focus on understanding. A natural framework for student study and learning arises: the mechanics of the cell
