 Everyone teaches something about basepairing in Introductory Biology, but sometimes it’s just in passing, with no greater emphasis than the diversity of sugar variants receives. But basepairing is the wellspring of life itself and lies at the heart of each of the major informational transactions (DNA replication, transcription, translation) as well as a diversity of more recently-discovered processes in regulation. More and more, we’re also seeing it as the great keystone of new therapies. What are the core concepts and how can we teach basepairing in order to showcase all of these aspects?
Everyone teaches something about basepairing in Introductory Biology, but sometimes it’s just in passing, with no greater emphasis than the diversity of sugar variants receives. But basepairing is the wellspring of life itself and lies at the heart of each of the major informational transactions (DNA replication, transcription, translation) as well as a diversity of more recently-discovered processes in regulation. More and more, we’re also seeing it as the great keystone of new therapies. What are the core concepts and how can we teach basepairing in order to showcase all of these aspects?
Monthly Archives: December 2014
Biological timers GTPases in translation, microtubules, signals
The main point in teaching with themes is to give students a meaningful sense of familiarity so that their travels from topic to topic are visits with friends and known quantities rather than onslaughts of fresh vocabulary and new un-anchored idea-pieces. One triplet that I have enjoyed using in introductory biology is the set combining 1) EF-Tu and its role in promoting accuracy in translation, 2) the GTPase timer in tubular that gives rise to the grow-catastrophe-disassemble cycle in microtubules, and 3) the ‘self shut-off’ timer features of many G-protein coupled receptors. These are evolutionarily related machines and each applies the concept of ‘timing’ to a different biological circumstance. Thus, besides providing three examples of how biological machines solve problems, this theme ties in to previous ones on ATPase (GTPase mechanism is identical; the attached base has nothing to do with catalysis) and evolution through duplication and modification.
What’s important about… Concepts in Translation
Teaching biology at the cell/molecular level is challenging; as the Bob Seger song puts it ‘Deadlines and commitments/what to leave in… what to leave out.” How to decide? The glib answer is “identify the key concepts.” A definition of these might be “what are the aspects and ideas without which the process would not make sense, and which, if changed, would rock the cell’s world.” So… what are the Big Concepts of translation? Starting with negation, these cannot be the names or even the structures of the twenty amino acids (believed to be a combination of what was on hand in abundance ‘in the beginning’ and ‘discoveries’ that were biochemically accessible) nor their codon assignments, which, may reflect some physical correspondences, but seem to largely be a ‘frozen accident’ that we should not expect to find elsewhere. Rant: it cannot be the ability to ‘translate’ a message in a codons-to-amino-acids look-up table, as this has nothing whatever to do with the biological process! And the core cannot be a laundry list of actors and factors.
The big concepts in translation…
Tatham Puzzle Collection: problem solving strategies
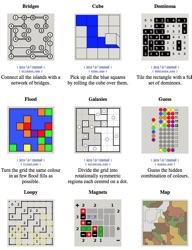 To my mind, there are two phases to a lot of classical ‘logic puzzles’. Take Sudoku (spoiler alert! Problem solving strategies already developed & revealed 🙁 ), for example. There is an initial phase where you figure out how to win; then you can play an infinite number of ‘success oriented’ games where you implement the strategies you developed initially, but rarely or never develop more. I think the initial phase holds the rich opportunities for giving folks an opportunity to develop their problem-solving skills in a meta-cognitive way. So, alongside Petals around the Rose, Bongard Problems, and PatternMaster, I’m linking to Simon Tatham’s Puzzle Collection because it hosts a variety of different problems. Not with the goal of allowing everyone an opportunity to play the one they like, but to explore and learn the ‘tricks’ of some that are unfamiliar.
To my mind, there are two phases to a lot of classical ‘logic puzzles’. Take Sudoku (spoiler alert! Problem solving strategies already developed & revealed 🙁 ), for example. There is an initial phase where you figure out how to win; then you can play an infinite number of ‘success oriented’ games where you implement the strategies you developed initially, but rarely or never develop more. I think the initial phase holds the rich opportunities for giving folks an opportunity to develop their problem-solving skills in a meta-cognitive way. So, alongside Petals around the Rose, Bongard Problems, and PatternMaster, I’m linking to Simon Tatham’s Puzzle Collection because it hosts a variety of different problems. Not with the goal of allowing everyone an opportunity to play the one they like, but to explore and learn the ‘tricks’ of some that are unfamiliar.
Bongard Problems: Quickie Scientific Method
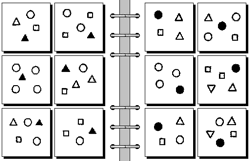 Bongard problems are an interesting way to challenge students to make discoveries using problem solving methods. An example (from the site linked at the beginning of this post) is shown below (hit ‘More’). All the Bongard problems have the same underlying premise: the group of six image/object collections on the left are united by some principle that is present in none of the six collections on the right. While it’s easy and fun to simply play this as a game, it’s another example of ‘discovery science’ based puzzle activities a la Petals Around the Rose or PatternMaster.
Bongard problems are an interesting way to challenge students to make discoveries using problem solving methods. An example (from the site linked at the beginning of this post) is shown below (hit ‘More’). All the Bongard problems have the same underlying premise: the group of six image/object collections on the left are united by some principle that is present in none of the six collections on the right. While it’s easy and fun to simply play this as a game, it’s another example of ‘discovery science’ based puzzle activities a la Petals Around the Rose or PatternMaster.

