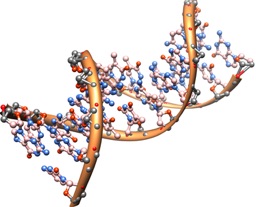
 Lists of the ‘characteristics of life’ are a common element in introductory biology early lectures. Generally, these focus on movement, energy conversion, organization, etc.–all legitimate concepts. But the role of self-assembly in biology must not be underestimated; it’s a key feature of the flow of information in the Central Dogma (through the specific partnering of bases), folding of proteins from linear strings of amino acids (readily specified by linear structure of nucleic acids) gives rise to the functioning machines at the heart of almost all cellular work and action, and even membranes, while not members of the Central Dogma club, have function that relies critically on aspects of self-assembly. Without these properties, life would not only be impossible–they’re prerequisites for life to evolve from non-life.
Lists of the ‘characteristics of life’ are a common element in introductory biology early lectures. Generally, these focus on movement, energy conversion, organization, etc.–all legitimate concepts. But the role of self-assembly in biology must not be underestimated; it’s a key feature of the flow of information in the Central Dogma (through the specific partnering of bases), folding of proteins from linear strings of amino acids (readily specified by linear structure of nucleic acids) gives rise to the functioning machines at the heart of almost all cellular work and action, and even membranes, while not members of the Central Dogma club, have function that relies critically on aspects of self-assembly. Without these properties, life would not only be impossible–they’re prerequisites for life to evolve from non-life.
Credit: This blog post was inspired by a recent discussion with my colleague Emily Dykstra, who besides countless general conversations with me on this topic, added membranes into the mix.
Overview: I think that if we’re seeking to teach concepts and understandings, start to finish our focus must be not shallowly on what happens, but the how and why. Since virtually all of the processes and pathways in Introductory Biology I (Cell/Molecular) involve molecules, ‘how they work’ is equivalent to ‘how they interact’ and ‘how decisions gate made’–be they decisions about whether to act, whether to stick, whatever. For a wide variety of biological processes these ‘decisions’ are automatic; when looked at in bunches we refer to the resulting processes as ‘self-assembly’. I’ve taken some liberties with the term in this post in order to make a point about how universal and foundational self-assembly in biology is.
Self-assembly in biology I: Genetic material; replication
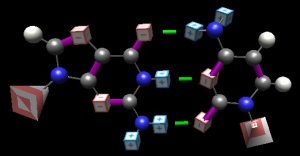 Before there were enzymes, before there were cells, before there was life, there were only chemicals and molecules. I’m not expert here, but my thinking is that initial cooperating entities had to ‘come together’ of their own accord… And of course, deep in the concept of ‘living’ is “able to reproduce.” For a nascent genetic material, this would entail the concept of getting copied without a copier, i.e. the machines would need to be self-copying. The only way this can happen is if an existing machine can more or less passively direct the creation of itself (or its complement, which in turn creates the original). The bases have this wondrous property–in a direct and meaningful sense, adenine ‘calls to’ thymine and guanine to cytosine without assistance and without need of a proofing mechanism: the basepairing face of guanine is complementary in a direct chemical way to that of cytosine.
Before there were enzymes, before there were cells, before there was life, there were only chemicals and molecules. I’m not expert here, but my thinking is that initial cooperating entities had to ‘come together’ of their own accord… And of course, deep in the concept of ‘living’ is “able to reproduce.” For a nascent genetic material, this would entail the concept of getting copied without a copier, i.e. the machines would need to be self-copying. The only way this can happen is if an existing machine can more or less passively direct the creation of itself (or its complement, which in turn creates the original). The bases have this wondrous property–in a direct and meaningful sense, adenine ‘calls to’ thymine and guanine to cytosine without assistance and without need of a proofing mechanism: the basepairing face of guanine is complementary in a direct chemical way to that of cytosine.
Of course, this capability is retained even today. DNA polymerase does not ‘pair bases’ as we too often carelessly say; it recognizes that the existing (template) base has ‘discovered’ a partner, and polymerase then ‘cements’ the relationship by performing the 3′ => 5′ backbone joining. Direct evidence that DNA polymerase has no ‘understanding’ of the Big 4 (A, G, C, T) comes from, for example, this demonstration by Benner and co-workers that DNA polymerase will cheerfully join ‘made up’ bases (ones with novel donor-acceptor patterns on their basepairing faces) with good fidelity. Even more dramatically, gifted with pre-activated substitutions instead of the usual XTPs, that bases can self-assemble into into templated polymers—protein-free RNA replication!
Self-assembly in biology II: Transcription
This really doesn’t require much, as transcription is essentially identical to replication with regard to the matching of nucleotide basis to the template. Again what really matters is that the bases match up with to their partners directly through chemical/electrostatic means, the machine doing the joining has no knowledge of legitimate versus illegitimate bases, and does not employ a”look up table” to determine which goes with which.
Self-assembly in biology III: Translation
Detailed in a previous post, there are myriad instances of base pairing and therefore self-assembly in the work of the ribosome. From the assembly of the small right ribosomal subunit with mRNA and initiator tRNA to the matching of each tRNA to the mRNA during elongation, the process matches closely with replication and transcription in that matching is passive, recognition and bond making is where the energy goes and the enzymes are involved.
Self-assembly in biology IV: Protein Folding
This one is huge, of course and is where much of our recognition of self-assembly or folding takes place. It can most readily be summarized by the”who will guard the guardians” concept: if proteins did not self assemble then presumably each would require a unique assembler corresponding to its unique final structure. Unless those assemblers are magical they in turn are proteins, each requiring its own assembler and so on and so on and so on.
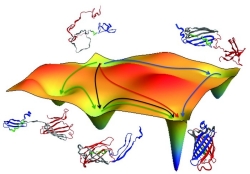
The path out of the wilderness of infinite regress is elegant and simple. Strings of amino acids are functionally ‘property beads’ on strings that have been selected through the long march of evolution to to reliably bring about unique three-dimensional structures: simply add water. In some ways, this is the central mystery of the central dogma: The linear transfer of information from a nucleotide-based language to an amino acid based-language buys us nothing unless that linear string of amino acids proceeds to become a functioning machine within the cell. Once again, the only way for such a machine to evolve is for that assembly process to be innate within each string of amino acids.
Try this link for a fun protein folding visualization (i.e., computer-based simulation) of GFP folding. Requires Flash plug-in. Both the image and the movie are derived from this PNAS article.
Self-assembly in biology V: Membranes
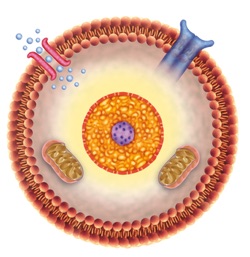
To the best of my knowledge, the debate is currently raging as to whether cells came first, surfaces came first, or informational/functional molecules came first. Regardless of order or simultaneity, the fact that lipid dropped into water spontaneously form sheets, micelles, and vesicles clearly paved the way for cellularity

Pingback: What’s important about… biology’s Central Dogma? – Stuff Bruce Says